A revolutionary discovery by researchers in Singapore and MIT has uncovered a new bacterial stress signalling system. This potentially opens doors to new methods of combatting antibiotic resistance, a growing public health concern.
The research team identified an enzyme, RlmN, that rapidly signals for the production of proteins allowing bacteria to adapt and survive. This could help design drugs that prevent this survival response, effectively tackling antibiotic resistance.
The Method Behind the Discovery
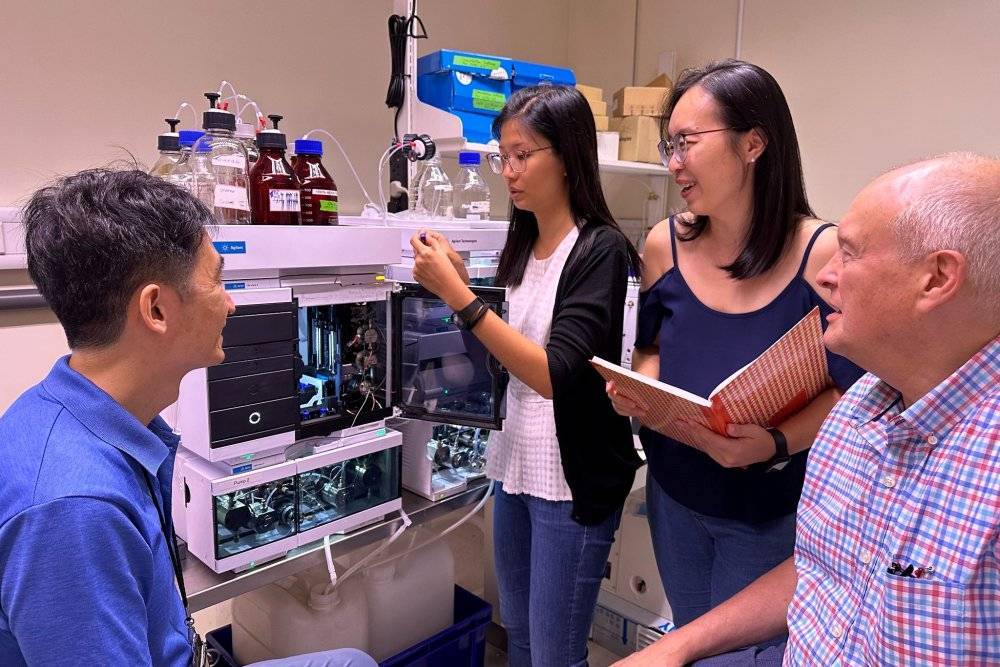
Using state-of-the-art mass spectrometry technology, researchers exposed E. faecalis cells, a common gut bacteria, to various antibiotics and toxic chemicals. Additionally, they identified the exact chemical changes that allow bacteria to survive antibiotic exposure.
A Widespread Phenomenon
The discovery of E. faecalis could also be present in other types of bacteria. Dr Lee Wei Lin, principal research scientist at SMART AMR, from the research team stated, “RlmN is not exclusive to E. faecalis but is widespread across bacteria. This emphasises the importance of this survival mechanism against antibiotics and host immune cells, and suggests that targeting RlmN could be a promising approach for developing new antimicrobial strategies.” The findings could be pivotal in developing new antimicrobial strategies.
Implications for Global Health
This research highlights two new avenues for antibiotic development. The two methods are through reactive oxygen species sensing and response, and through targeting RNA modifications. The discovery could lead to the development of new classes of antibiotics, something not seen for decades.
Dr Megan McBee, Senior Scientific Director at SMART AMR, stated that “This discovery can potentially open up two avenues for new targets/classes for antibiotic development, ie, through reactive oxygen species sensing and response, and through targeting RNA modifying enzymes. Antibiotics and host immune cells use reactive oxygen species to neutralise bacteria. Hence, reactive oxygen species play fundamental roles in shaping bacterial evolution. We believe that by understanding how bacteria deal with reactive oxygen species, we will be able to identify new targets for antibiotic development.”
A Silent Pandemic
Antimicrobial resistance is often referred to as a silent pandemic, reducing the efficacy of existing antibiotics and increasing mortality rates. Understanding these cellular adaptation mechanisms will pave the way for developing new therapies, a priority for global public health.
Conclusion
The groundbreaking research carried out by the Singapore-MIT Alliance for Research and Technology (SMART). Additionally, it is supported by the National Research Foundation Singapore. The study stands as a beacon of hope in the fight against the global threat of antibiotic resistance.
This pioneering discovery represents not just a scientific triumph, but a major advancement in global healthcare. By unlocking new pathways and understanding these intricate biological processes, researchers have brought us a step closer to a world free from the devastating effects of antimicrobial resistance. Therefore, the implications of this work are profound. This may well signal a turning point in the fight against one of the most challenging medical crises of our time.
The research is carried out by the Antimicrobial Resistance (AMR) Interdisciplinary Research Group (IRG) at Singapore-MIT Alliance for Research and Technology (SMART), in collaboration with Singapore Centre for Environmental Life Sciences Engineering (SCELSE), Nanyang Technological University Singapore (NTU Singapore) and Massachusetts Institute of Technology (MIT), and supported by the National Research Foundation (NRF) Singapore under its Campus for Research Excellence And Technological Enterprise (CREATE) programme.